 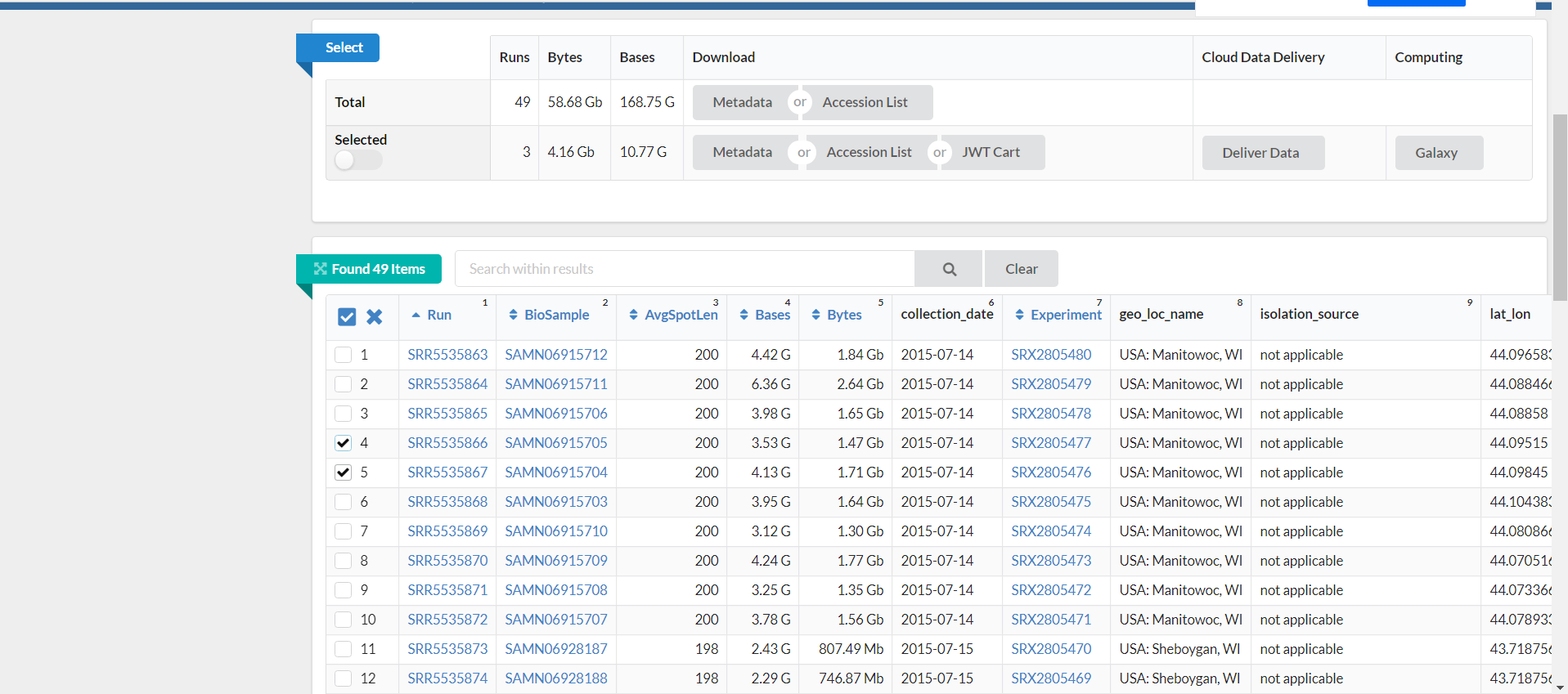
Write.table ( x= metadata], file = paste0 ( "./clean_datas/", i, "_SRR_Acc_List.txt" ), sep = " \t ", row.names = F, col.names = F, quote = F)įor (i in list.files ( "./clean_datas", pattern = "_SraRunTable. Write.table ( x= metadata], file = paste0 ( "./clean_datas/" ,i, "_SraRunTable.txt" ), row.names = F, sep = " \t ", col.names = T, quote = F) Write.table ( x= meta, file = paste0 ( "./clean_datas/", j, "_SraRunTable.txt" ), sep= " \t ", row.names = F, col.names = T, quote = F) Write.table ( x= vec, file = paste0 ( "./clean_datas/", j, "_SRR_Acc_List.txt" ), sep = " \t ", row.names = F, col.names = F, quote = F) Not_shotgun] % filter ( if_any ( everything (), ~ str_detect (., regex_match))) # tmp2 = str_split(i, pattern="_", simplify = T) Not_nb = matches ( vars= metadata], c ( "pcr", "Amplicon", "OTHER", "16S" ))Ĭat (tmp, " \n ", "column", paste0 ( colnames (metadata]), collapse = " | " ), " \n ", "seems to be only 16S rRNA gene \n " ) Nb = matches ( vars= metadata], c ( "shotgun", "WGS", "whole genome", "all_genome", "WXS", "METATRANSCRIPTOMIC", "NextSeq", "WholeGenomeShotgun" ))Ĭat (tmp, " \n ", "column", paste0 ( colnames (metadata]), collapse = " | " ), " \n ", "contains a pattern related to SHOTGUN sequencing, there is shotgun innit \n " ) #find out which accession_list contains shotgun data Setwd ( "/media/remy/Stockage/Database_fastq/SOP_Early_LIFE" ) SRA entrez usually covers for SRA, ENA, DDBJ and more. Then on the top right of the panel you’ll find “Send to” button. Paste the accession number in the searching zone, press enter. You need to go to the SRA entrez on NCBI and use the accession number to find your samples. txt format from the run selector or the sra entrez search. 
To download this tar file copy the below commands and paste it in your terminal and press enter. We will download tar.gz of sra toolkit for this. Go to SRA entrez in NCBI : įirst get the filenames in. Download SRA Toolkit Now you will have to download the sra toolkit inside this directory. Once the directory is set you can start to get your data. 
Users/mala76/sratoolkit/sratoolkit.2.10.9-mac64/bin/vdb-config -i So change this to a directory with a large storage capacity. If you don’t perform this step the toolkit will download in the directory were you put it, usually in your root. Rapidly you can just run this in your terminal and chose the directory you want the SRA toolkit to download into. You need to configure it as explained here : You first need to download the SRA toolkit from NCBI : The toolkit will download SRA formatted files and then transform them in fastq. How to install sra-toolkit ubuntu package on Ubuntu 20.04/Ubuntu 18.04/Ubuntu 19.04/Ubuntu 16.04 - Server Hosting Control Panel - Manage Your Servers. You will need to install, configure and get accession list for your samples. NCBI made a very useful tool to download quickly and accurately large sequencing data stored on SRA, ENA or DDBJ servers. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
